Even Mitochondrial RNA must “Grow Up”
Human beings must first learn to crawl and then walk before they can run. So too must RNA undergo maturation, involving several transformations, before it is able to play its part in the synthesis of proteins. Understanding the process of mRNA maturation in mitochondria is one of the focuses of the Hällberg lab.
While most of the proteins that work inside the mitochondrion, the power plant of the cell, are encoded in the nuclear genome, 13 proteins critical for power generation are encoded on the mitochondrion’s own genome called the mitochondrial DNA (mtDNA). Similar to normal DNA, mtDNA undergoes the process of transcription during which RNA is synthesized. This RNA is essentially still in its infancy and must undergo several changes before it is considered to be mature and can take part in translation: the synthesis of proteins.
Polyadenylation is one of these changes, during which poly(A) polymerase, an enzyme, builds a tail of adenine nucleotides onto the 3’-ends of mRNAs. “Polyadenylation alters the fate of mRNAs in several ways. It can increase mRNA stability, stimulate translation initiation, promote degradation or be required for completing certain stop codons that are not encoded in mtDNA,” explains CSSB Group leader Martin Hällberg “Polyadenylation is critical for mitochondrial translation and hence energy generation by the mitochondrion.”
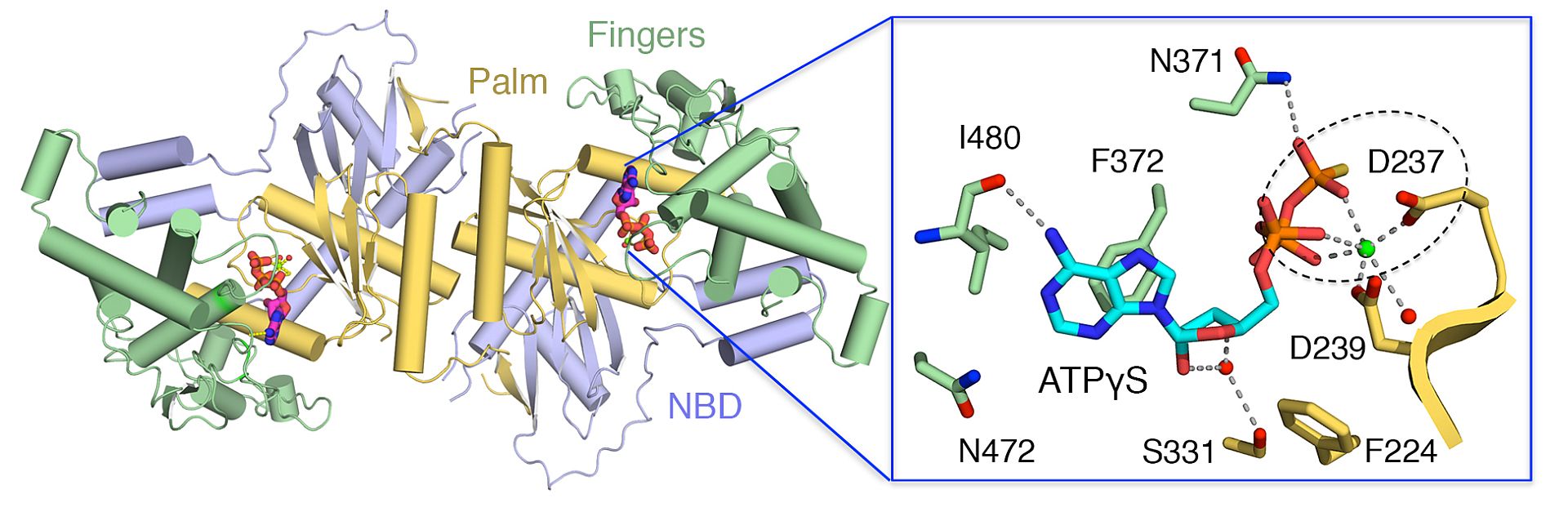
A recent paper published by Hällberg and his colleague, Mikalai Lapkouski, reveals several structures of the mitochondrial poly(A) polymerase (mtPAP) that adds polyadenines to the 3’-ends of mitochondrial mRNAs. Using the beamlines on the DESY Campus, Hallberg’s group analysed the crystal structures of chicken mtPAP, which is approximately 60% identical to that of human. The structures generated provide detailed and informative insight into the mechanisms of dimerization and ATP-specificity of mtPAP. mtPAP is, interestingly enough, only active as a dimer. “What we see is that only one of the two active sites is active at one single time,” explains Hällberg “the generation of heterodimeric enzyme in which one of the two monomers is catalytically inactive shows that the two active sites can operate independently.”
Understanding how mtPAP operates also shed some light on the causes of spastic ataxia 4 (SPAX4), a rare neurodegenerative disorder affecting balance, speech, movement of the arms, legs, and tongue. A missense mutation in human mtPAP was recently found to be responsible for this disorder. “This mutation diminishes mtPAP’s polymerization activity, which results in shorter poly(A) tails in mRNA and ultimately dysregulates the post-transcriptional expression of proteins.” explains Hällberg. The structures revealed by the Hällberg lab provide a structural understanding of the enzymatic defect that results from the SPAX4 missense mutation.
“We are now trying to get a complex with an actual mRNA substrate bound to understand how the substrate and enzyme interact. In addition, this will provide even more insights into the molecular basis of the SPAX4 mutant,” says Hällberg of his group’s upcoming plans.



