Research Projects
Gamma-Secretase in Alzheimer Disease
γ- secretase is a membrane protease complex which consists of four subunits: presenilin (PS), nicastrin (NCT), APH-1 and PEN-2 . γ-Secretase cleaves over 90 type-I integral membrane proteins which are involved in many physiological and pathological process.
Uncleaved Presenilin functions als as a Ca-channel whereby it may affect pathologies. Furthermore it is the receptor for papilloma virus infection.
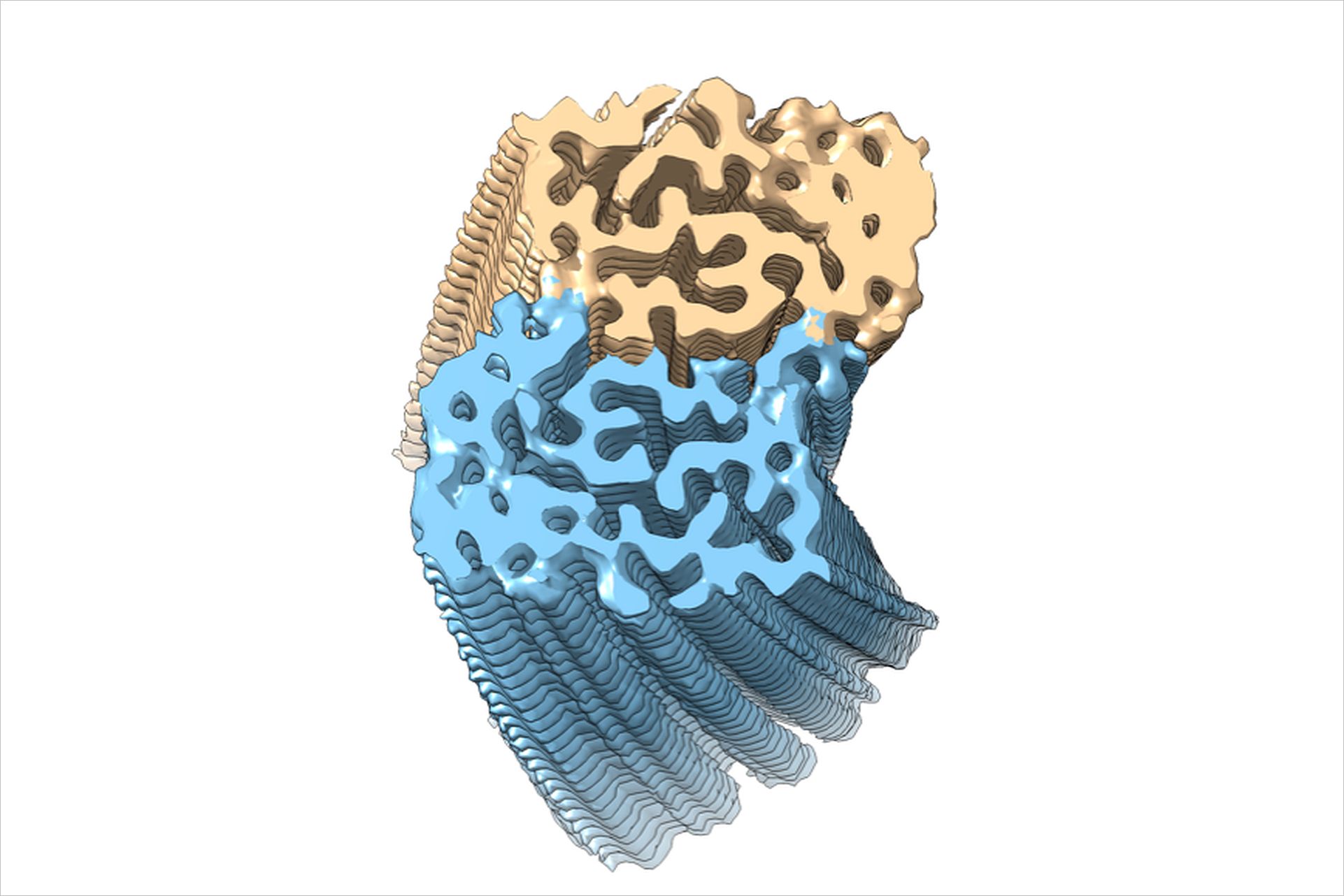
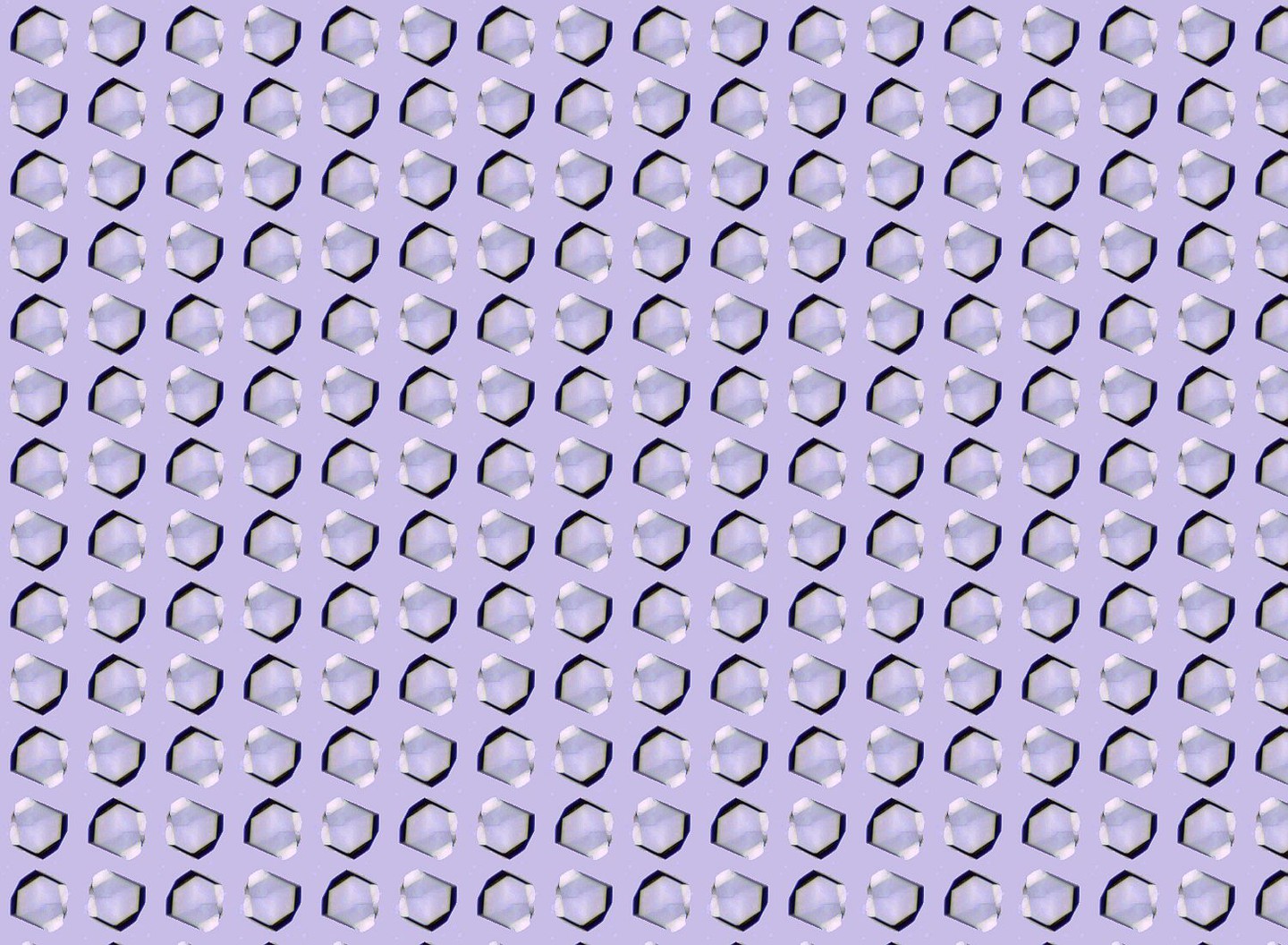
The most widely studied γ-secretase substrates are Notch and C-terminal fragments of β-amyloid precursor protein (APP-CTFs). The cleavage of Notch by γ-secretase releases the intracellular domain to the nucleus, which in turn affects the development of cells and cancer. The cleavage of APP-C99 by γ-secretase generates amyloid-beta peptides (Aβ) of varying lengths, among which the longer peptides -particularly Aβ42 and Aβ43 - form fibrils (s.Fig.1) and are the key components of Aβ plaques, a hallmark of Alzheimer's disease (AD).
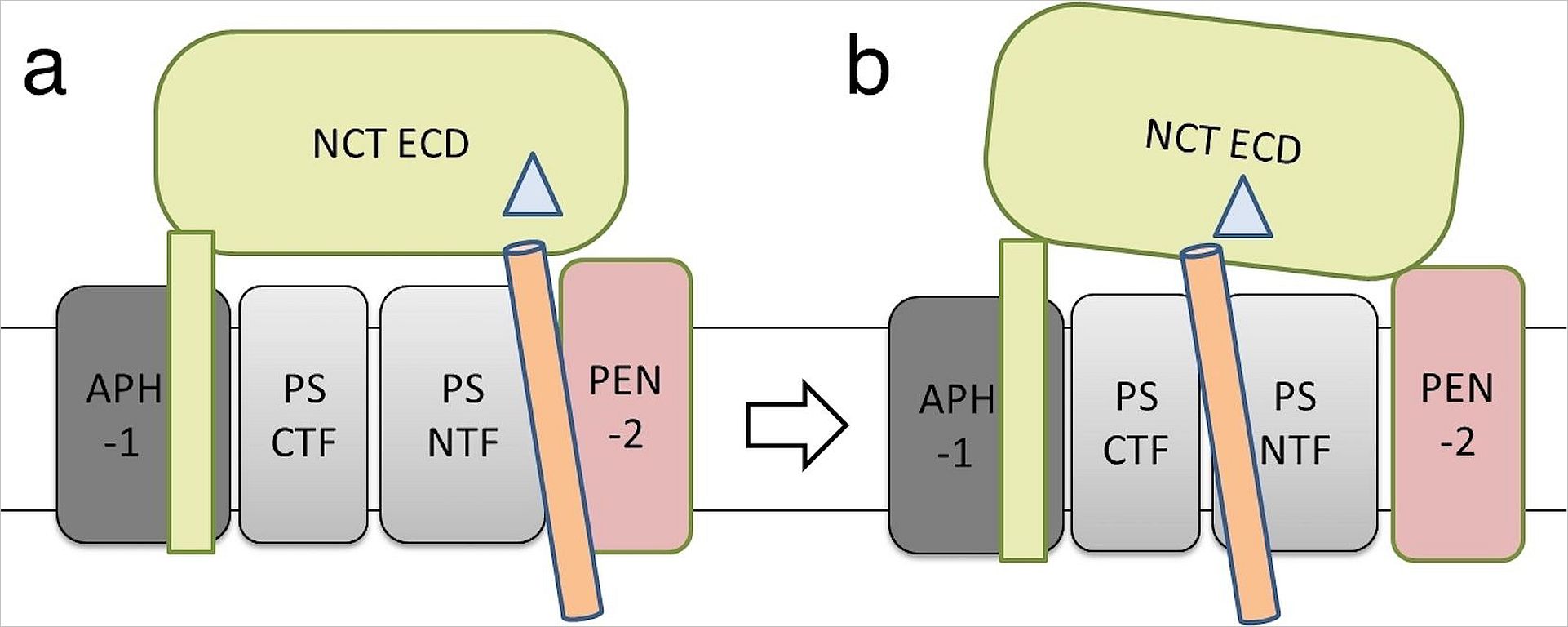
We found evidence that NCT (yellow) may is part of an initial substrate docking site(s. Fig. 2) from where the substrate (orange) needs to be translocated to the activated catalytic subunit PS (grey), where the substrate is processed. The role of APH-1 (dark grey) and PEN-2 (pink) in complex assembly and catalysis is obscure. The pathological change of the product spectrum may be due to a disturbance of substrate translocation.
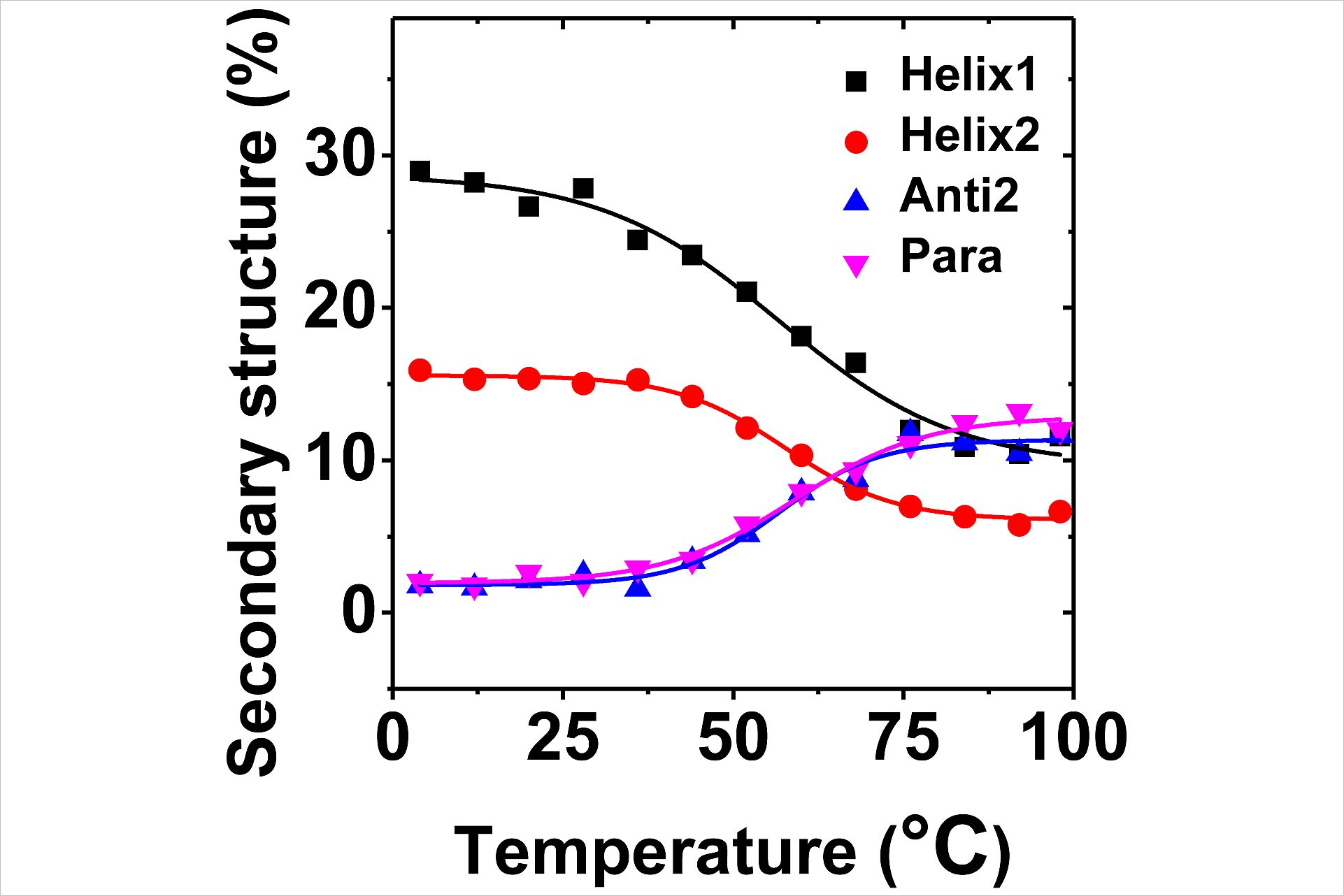
The deconvoluted component spectra of presenilin showed a thermally inducible structural transition from helix to strand (s. Fig. 3). Half of the helical content restructured. Whereas the content of unordered structure and turns increased only slightly, more than ~60% of helical structure turned into β-structure. The linear dependence and the presence of an isodichroic point are evidence for the existence of two different secondary structures of the protein: A low temperature form rich in α-helices and a high-temperature form rich in β-structures. This suggests that the AD related catalytic dysfunction of γ- secretase may be related to structural changes that normally occur only to a small amount at body temperature, but are elevated for mutated Presenilins as found in the cases of familial (inherited) Alzheimer disease.
Par4: Induction of Apoptosis in cancer cells
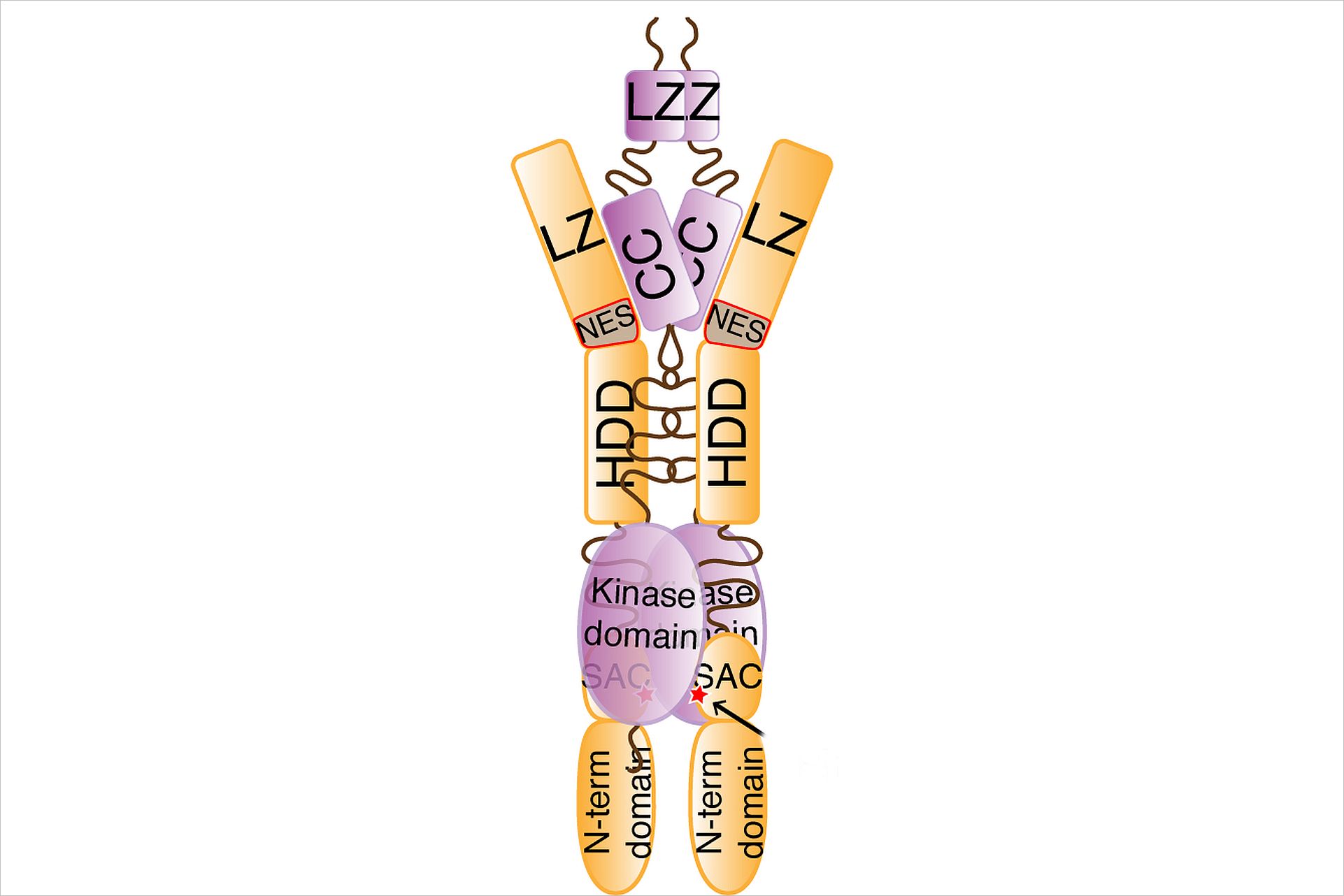
Par-4, the prostate apoptosis response factor 4 (aka Pawr) is a unique pro-apoptotic protein with the ability to induce apoptosis selectively in cancer cells and is involved in HIV-encephalitis. The X-ray crystal structure of the regulatory domain of Par-4 (Par-4CC) showed that Par-4 homodimerizes by forming a parallel coiled-coil structure. The N-terminal half of Par-4CC contains the homodimerization subdomain. This structure includes a nuclear export signal (Par-4NES) sequence, which is masked upon dimerization indicating a potential mechanism for nuclear localization. The heteromeric interaction models specifically showed that charge interaction is an important factor in stability of heteromers of the C-terminal leucine zipper subdomain of Par-4 (Par-4LZ). These heteromer models also displayed NES masking capacity and therefore the ability to influence intracellular localization. An important factor in regulation of the apoptotic function is the formation of a complex with Dapk3 (death associated protein like kinase). The structural information from Par4 allows the formulation of a first model of this complex (s. Fig) and an understanding of the role of the functional domains and subdomains in the regulation of apoptosis.
Membrane Protein Methods
The investigation of membrane proteins in general and mammalian in particular poses substantial problems in obtaining the proteins in sufficient quantity and quality to successfully investigate the mechanisms of their molecular pathology. We developed a lipidic-cubic phase crystallization approach which allows controlling the structure of the meso-phase which provides the crystallization matrix and thereby steering crystallization.
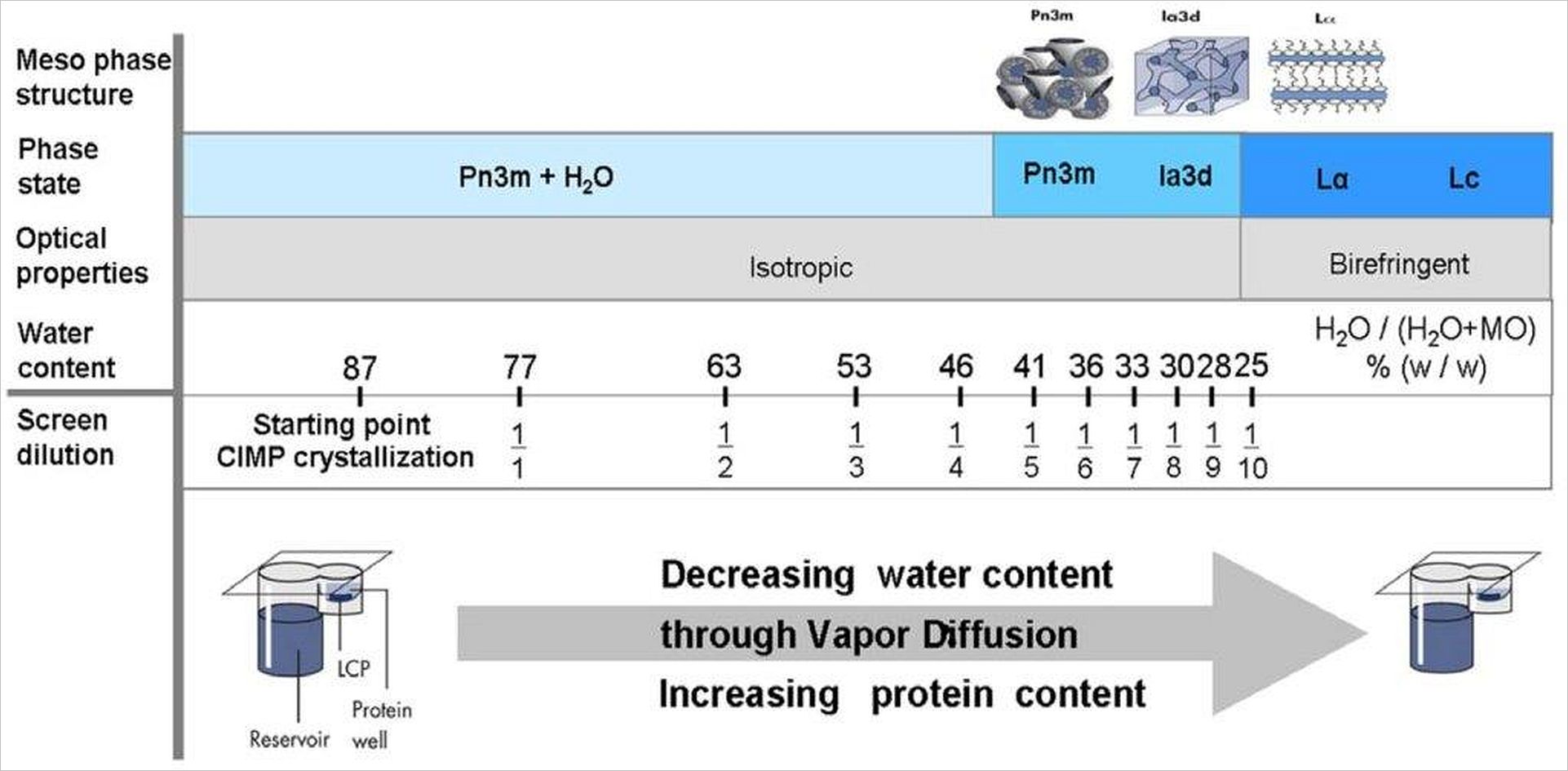
The lipidic meso-phases (Pn3m, Ia3d, Lα, Lc) are formed by monoolein with different amounts of water. The formation of these crystalline phases can be monitored by change of optical properties. Phase formation in the presence of membrane protein leads to insertion of protein into the hydrophobic layers of the monoolein. In case of the cubic phases (Pn3m, Ia3d) 3-dimensional diffusion of the protein within the membrane-like layer is possible: The crystalline meso-phase acts as a solvent for the membrane protein. Reducing the water content of meso-phase by vapor diffusion induces crystallization (phase separation) of the membrane protein. J. Kubicek, et al. PlosOne 7(4): e35458 (2012).
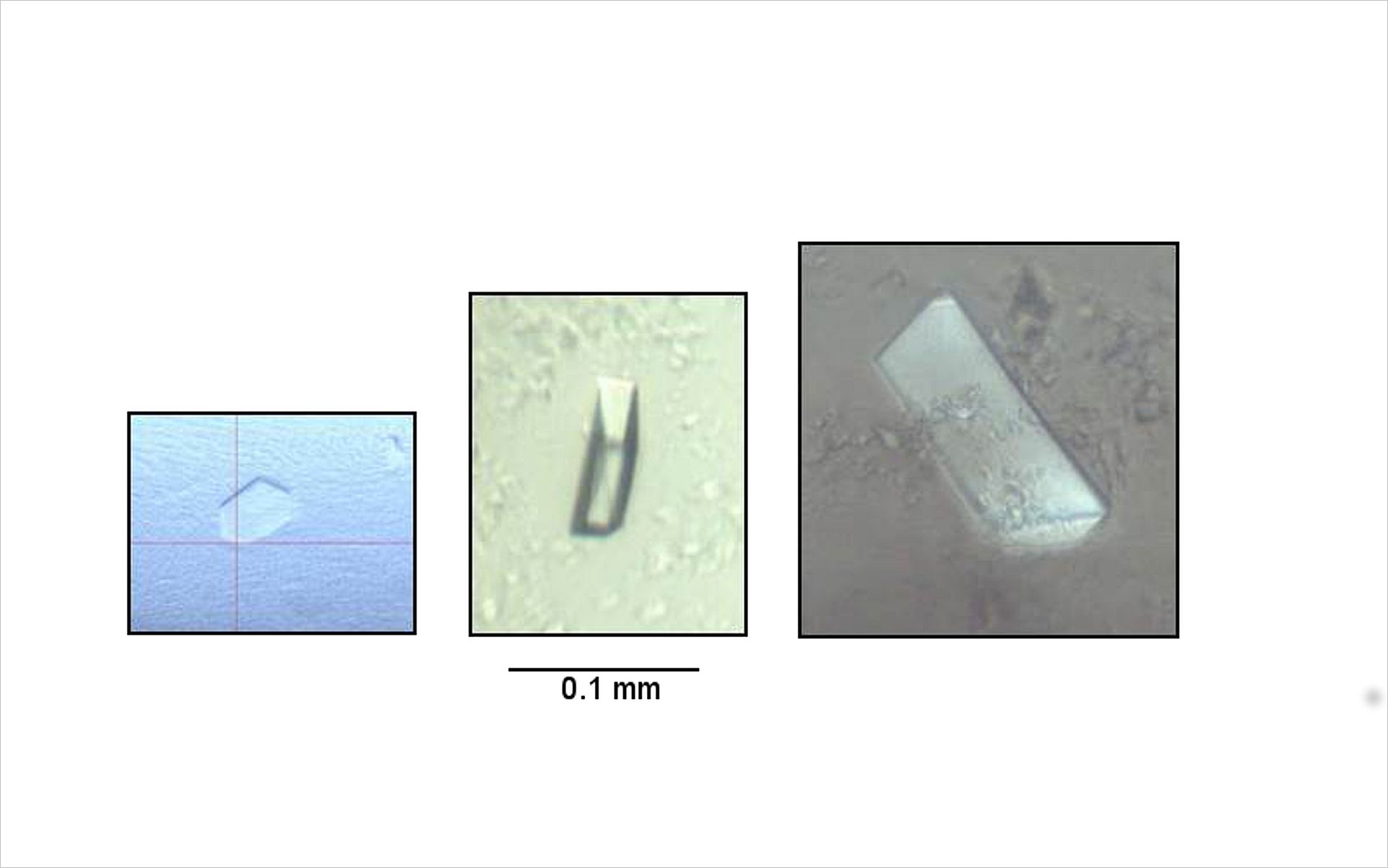

One of the major obstacles in the structural analysis of human membrane proteins is the optimization from a 1st-hit (left) to diffracting crystals (right). If properly optimized crystallization of high quality crystals can be achieved within half a day (see Video)
The need to solubilize membrane proteins for investigation poses another major problem because detergents can disrupt the protein structure and functionality. An interesting alternative strategy is the solubilisation of membrane proteins by genetic engineering. 20 to 25 % of hydrophobic amino acids at the protein surface need to be mutated to more soluble ones to transform the insoluble membrane protein into a soluble. There is a specific exchange code for aminoacids that minimizes functional and structural disturbances: the QTY code (S. Zhang et al, PNAS, 115 (37) E8652-E8659 (2018).
FUTURE PROJECTS AND GOALS
In collaboration with the Kolbe group we plan to establish at CSSB a platform for the analysis of infection factors of Pseudomonas aeruginosa. Pseudomonas aeruginosa is an adaptive environmental bacterium and an important opportunistic pathogen, which causes devastating acute as well as chronic, persistent infections. Its ecological success can be attributed to the dynamic expression of interacting regulatory networks that drive bacterial adaptation mechanisms and the expression of virulence traits. Our aim is the systematic structural and functional characterization of putative virulence proteins from uncharacterized open reading frames in P. aeruginosa.

