Structure of Type VII Secretion System Revealed
In a new study, published in Nature, CSSB researchers and international collaborators reveal new insights into the structure of type VII secretion systems from Mycobacterium tuberculosis. Type VII secretion systems are molecular machines that play key roles in the infection cycle of many pathogenic mycobacteria, including the notorious Mycobacterium tuberculosis. A deeper understanding of the structure and function of these systems can enable the development of novel therapies for the treatment of tuberculosis.
Prior to the corona virus pandemic, tuberculosis was the leading cause of death worldwide from a single infectious agent. Mistakenly considered by many to be a disease of the past, tuberculosis is a disease that still kills four thousand people every day (approximately 1.4 million every year) (1). With the spotlight placed on combatting COVID-19, the fight against tuberculosis has now suffered additional difficulties, due to decreased options for testing and treatment during lockdown periods. WHO models are currently estimating an excess of half a million TB deaths in 2020 and a decade-long setback in the fight against TB (2).
To initiate infection, Mycobacterium tuberculosis needs to transport bacterial proteins across its distinct, impermeable double-membrane cell envelope. “The pathogen possesses an arsenal of dedicated molecular machineries, called type VII secretion systems (T7SS), that facilitate this transport,” explains Wilbert Bitter from the Vrije Universiteit Amsterdam/Amsterdam UMC “and this pathogen has five different types of these systems that are not only central for virulence but also critical for nutrient uptake which makes ideal drug targets to fight the disease.”
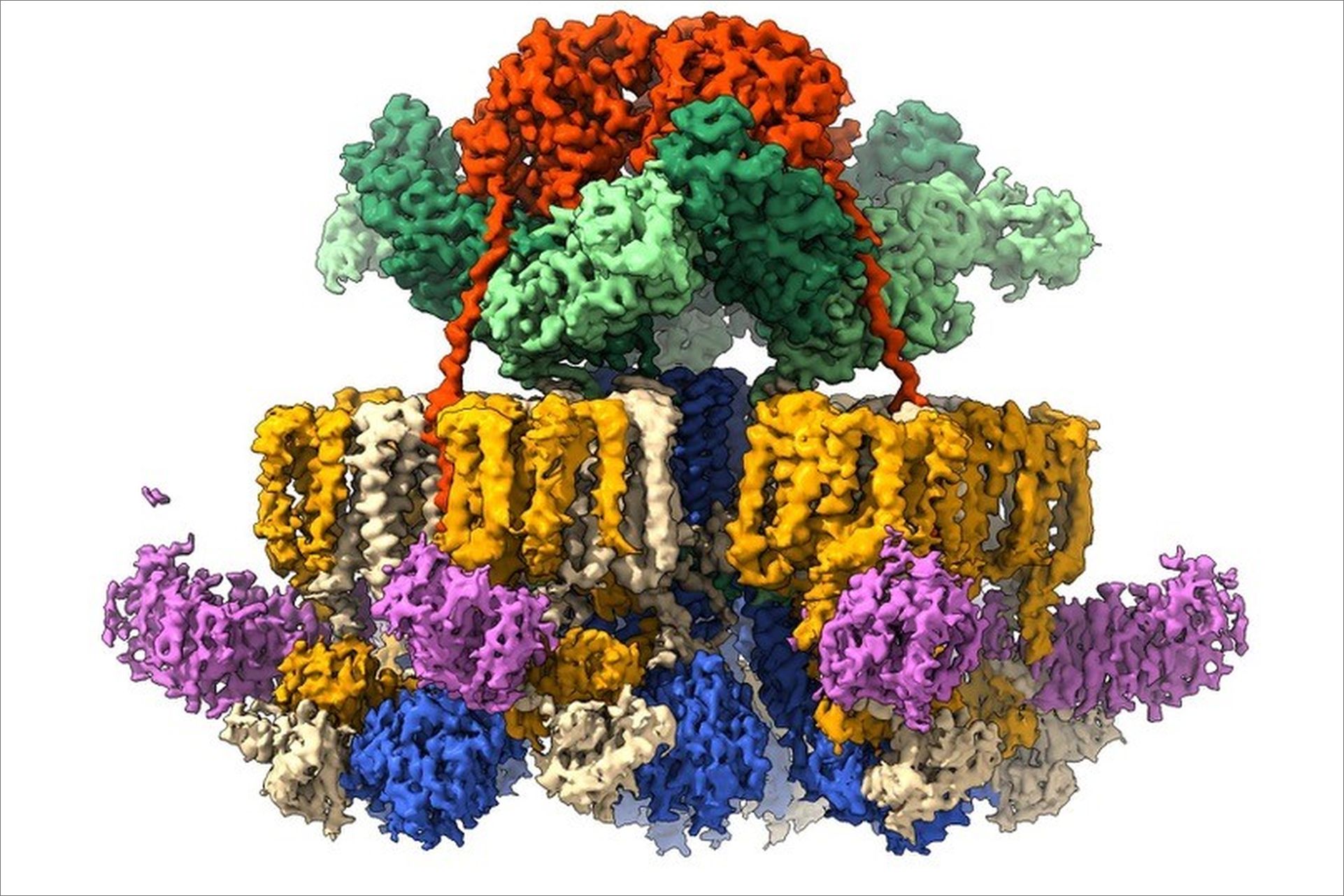
While previous studies conducted by the researchers revealed that four T7SS components form a six-sided star-shaped complex, high-resolution structural information and mechanistic insights could not be determined due to the instability of the complex (3). “We now understand how the components are kept together as one stable assembly, allowing us to see that this large molecular machine has a staggering 165 membrane anchors”, notes investigator Edith Houben from the Vrije Universiteit Amsterdam.
To bypass the high infectivity of pathogenic mycobacteria, Amsterdam UMC researcher Roy Ummels reconstituted one of the nanomachineries of Mycobacterium tuberculosis in non-pathogenic mycobacterial species. This allowed the purifications of large quantities of the assembly for cryo-EM analysis. After many trials to obtain a sample that is suitable for cryo-EM imaging, high-resolution information was collected at CSSB’s cryo-electron microscopy facility. “It was a very exciting journey to see how the quality of the sample has improved over the years to the point where we were impatient for the microscope to be available to us” notes CSSB researcher Jiri Wald.
The high-resolution structure not only confirmed the previously seen star-like shape of the assembly, but also revealed how this complex is stabilized by an additional fifth component, the MycP protease, which is essential for T7SS functioning. The structural model shows a group of three MycP enzymes capping a dome-like chamber at the top of the complex. Each MycP sits on top, like an inverted cherry, and provides stability by increasing the number of contact points among subunits of the complex. “In the absence of MycP the entire complex becomes more flexible, and the arms of the star begin to wobble”, explains CSSB researcher and first author Catalin Bunduc “The complex with MycP is like a stable suspension bridge and when the suspenders (MycP) are removed, the complex lacks support and sways like a wobbly footbridge.”
The data also provided new insights into how T7SSs could be able to secrete these proteins in a controlled manner without allowing other molecules to leak in and out. Proteins are usually transported in an unfolded state, meaning that prior to traveling through a secretion channel the protein is untangled into a chain-like structure, reducing its width. T7SS, however, secretes proteins in at least a partially folded manner thus requiring a relatively large opening in the bacterial membrane. “The membrane anchors of the motor protein assemble into bundles that seal the central secretion channel” notes CSSB researcher Dirk Fahrenkamp, “And the machinery seems to form two communicating chambers, one on each side of the inner membrane, potentially preventing leakage when the pore is opened."
While tuberculosis can be treated with antibiotics, drug resistant strains are becoming increasingly more prevalent and in 2019 these represented 3.5% of new TB cases and 17.7% of previously treated cases. The structural insights gained by the researchers provide a starting platform for the identification of domains and interactions that could be targets for drug development. “Our results are a huge step forward in understanding T7SSs and Mycobacterium tuberculosis itself” explain the authors “we can now investigate potential drug binding sites that inhibit the function of T7SS and prevent the propagation of infection.”
“This breakthrough not only reveals new possibilities in the fight against tuberculosis but also emphasizes the societal importance of fundamental research. Breakthroughs, like this one, can ultimately lead to practical applications that save lives,” marks CSSB investigator Thomas Marlovits.
Source:
Catalin M. Bunduc, Dirk Fahrenkamp, Jiri Wald, Roy Ummels, Wilbert Bitter, Edith N. G. Houben & Thomas C. Marlovits: Structure and dynamics of a mycobacterial type VII secretion system; Nature 2021; DOI: 10.1038/s41586-021-03517-z
DESY News Story:
https://www.desy.de/news/news_search/index_eng.html?openDirectAnchor=2068&two_columns=0
References:
(1) WHO, Global tuberculosis report (2020).
(2) WHO, Impact of the COVID-19 pandemic on TB detection and mortality in 2020 (2021).
(3) Beckham, K. S., Ciccarelli, L., Bunduc, C. M., et al. Structure of the mycobacterial ESX-5 type VII secretion system membrane complex by single-particle analysis. Nat Microbiol 2, 17047, (2017).